Puzzle-GEM User Manual
Overview
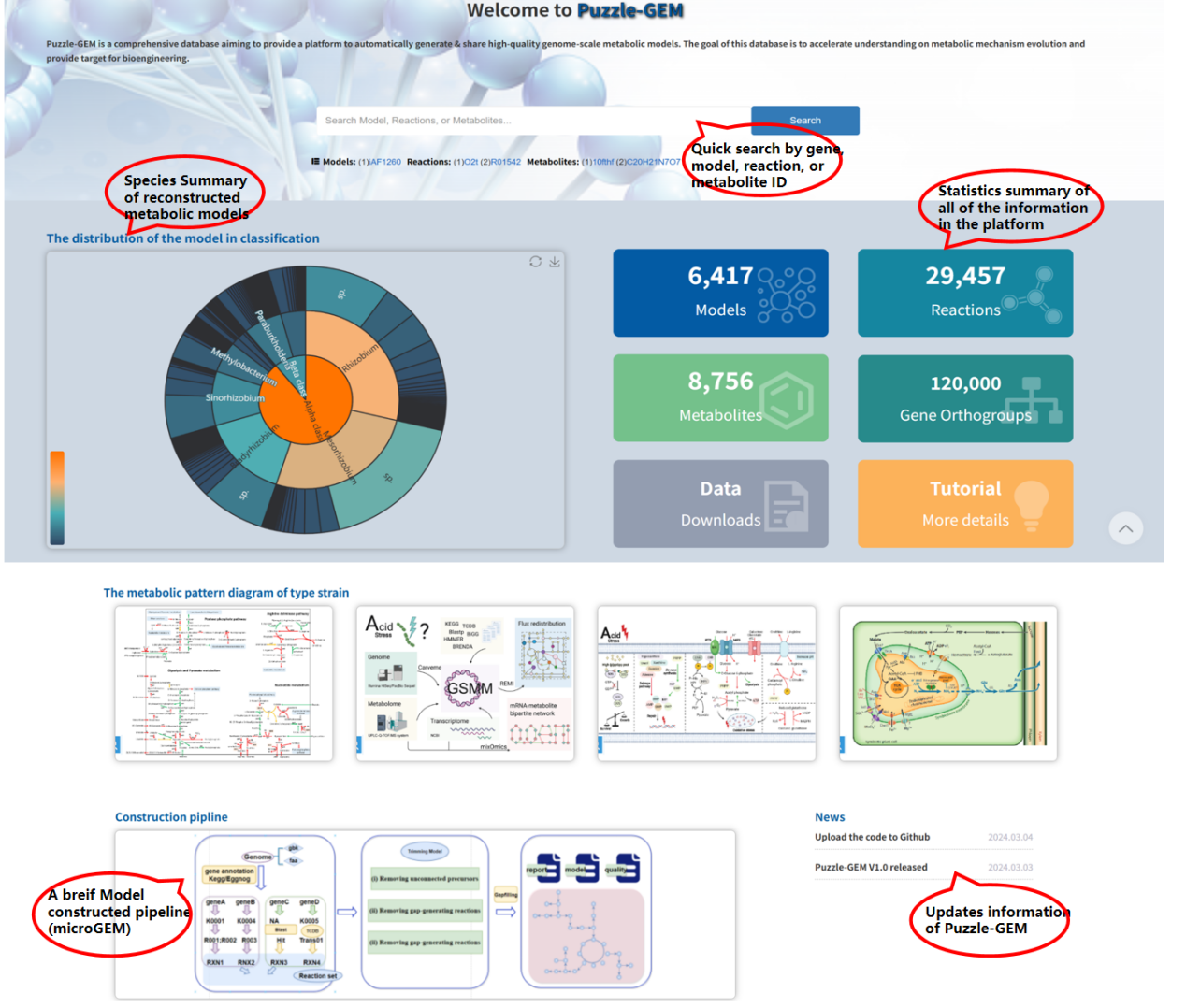
Home
Browse
Browse Models
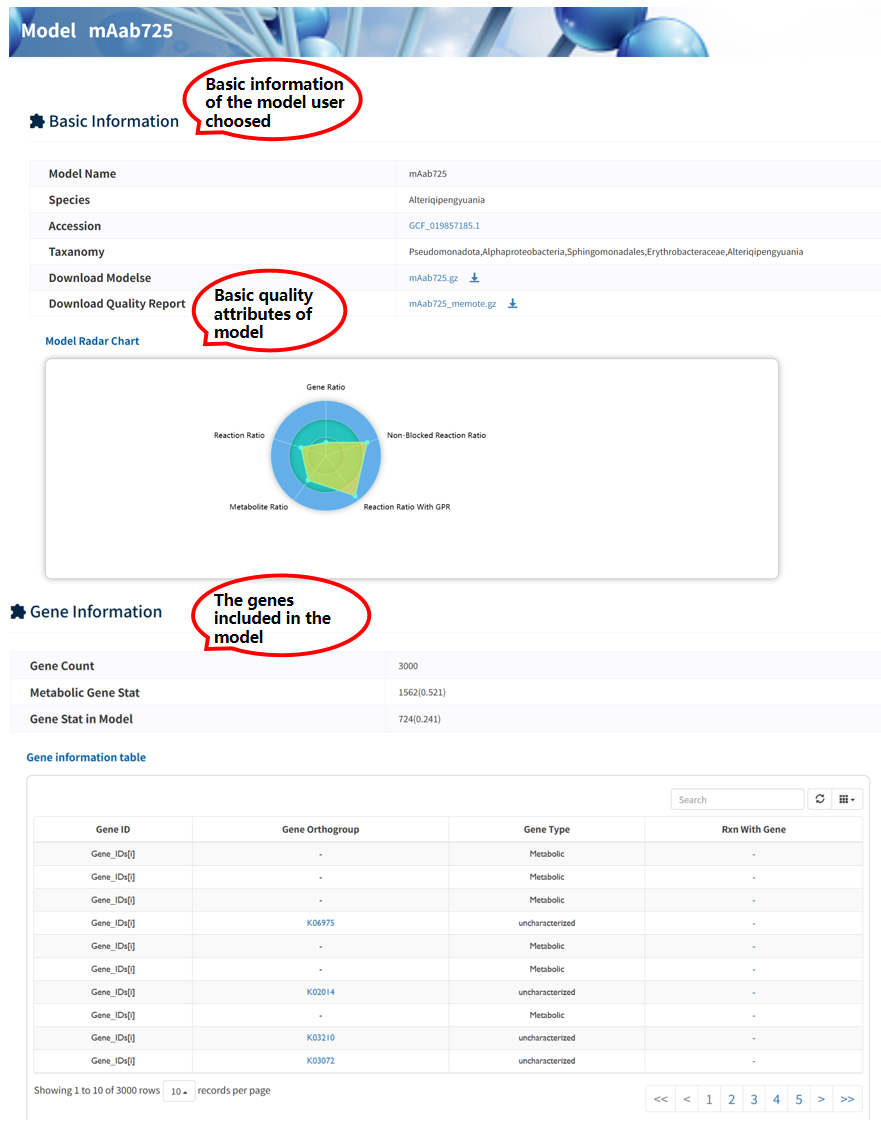
The Browse Models page provides summary statistics and key features of all GEMs, including species information, reaction, metabolite, and gene counts, as well as access to model files and quality reports.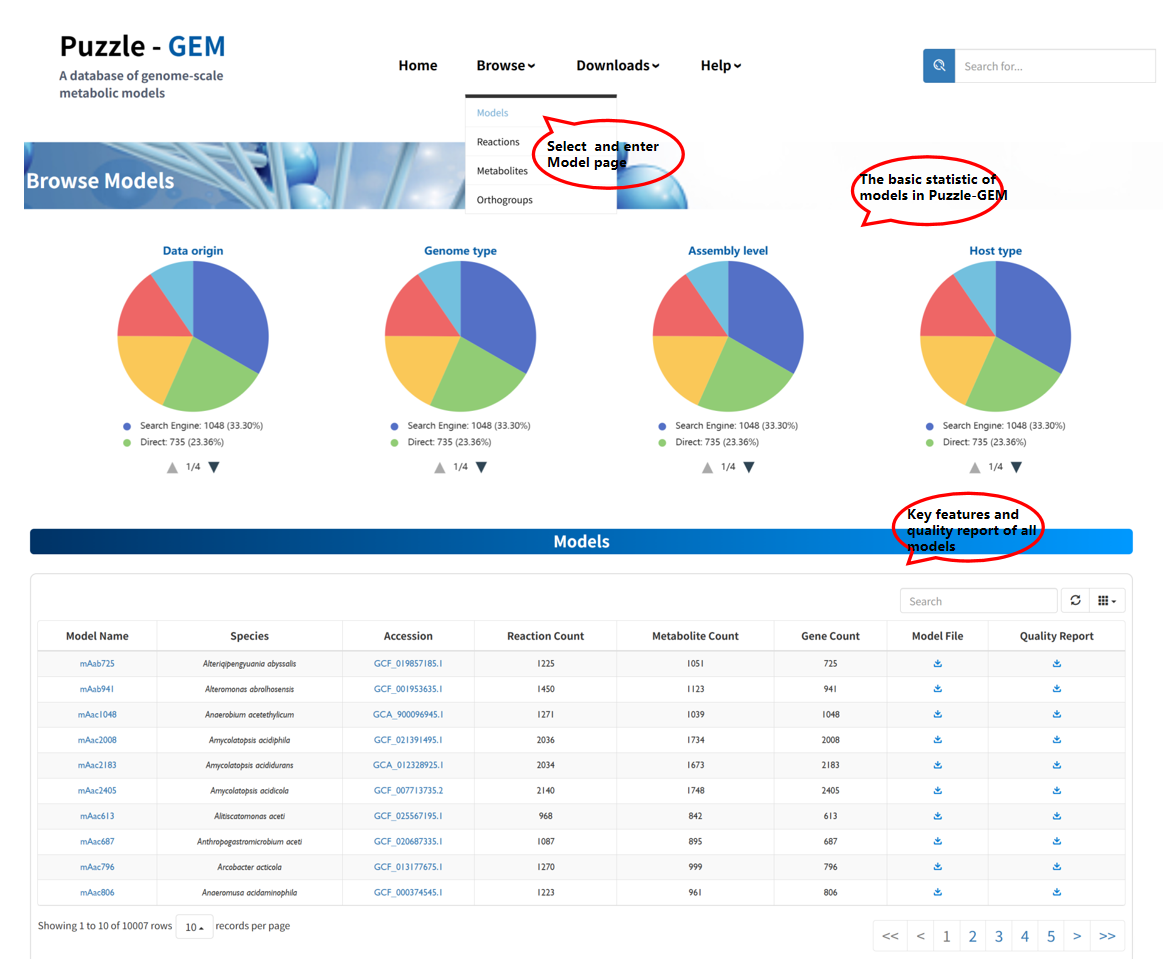 Users can download the model file and its quality report file. In addition, clicking the accession number redirects to the corresponding genome entry in NCBI, while clicking the model name navigates to a detailed model information page.
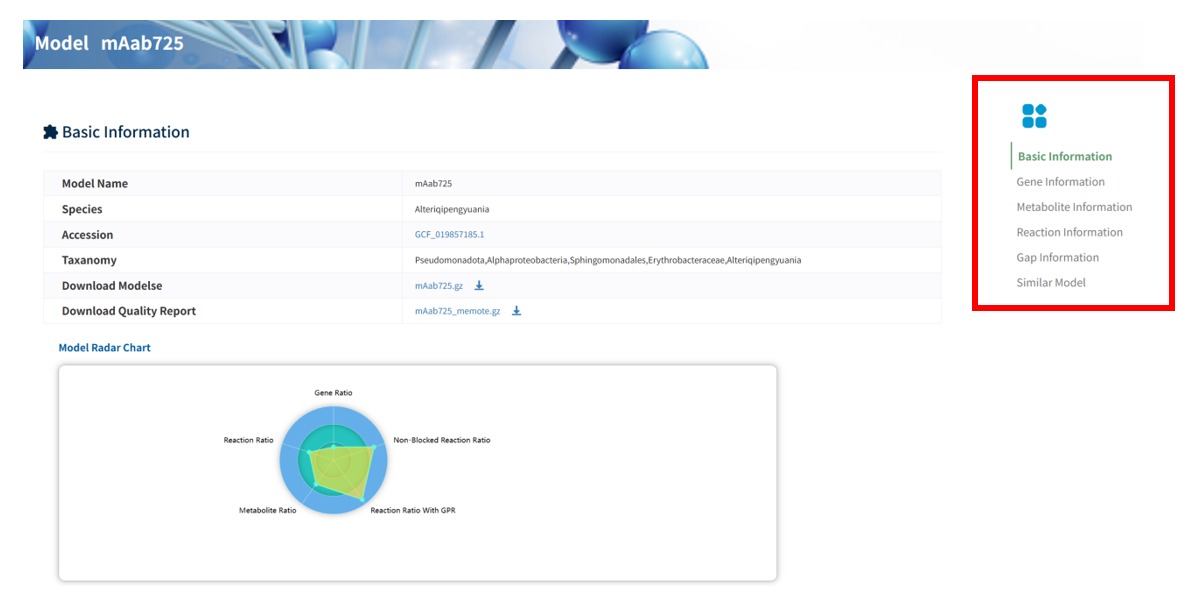
Each reconstructed GEM is associated with a dedicated Model information page, where comprehensive details of the model are provided. A navigation panel on the right allows users to quickly access different sections, including 6 sections: Basic Information, Gene Information, Metabolite Information, Reaction Information, Gap Information, and Similar Models.
Users can download the model file and its quality report file. In addition, clicking the accession number redirects to the corresponding genome entry in NCBI, while clicking the model name navigates to a detailed model information page.
Each reconstructed GEM is associated with a dedicated Model information page, where comprehensive details of the model are provided. A navigation panel on the right allows users to quickly access different sections, including 6 sections: Basic Information, Gene Information, Metabolite Information, Reaction Information, Gap Information, and Similar Models.
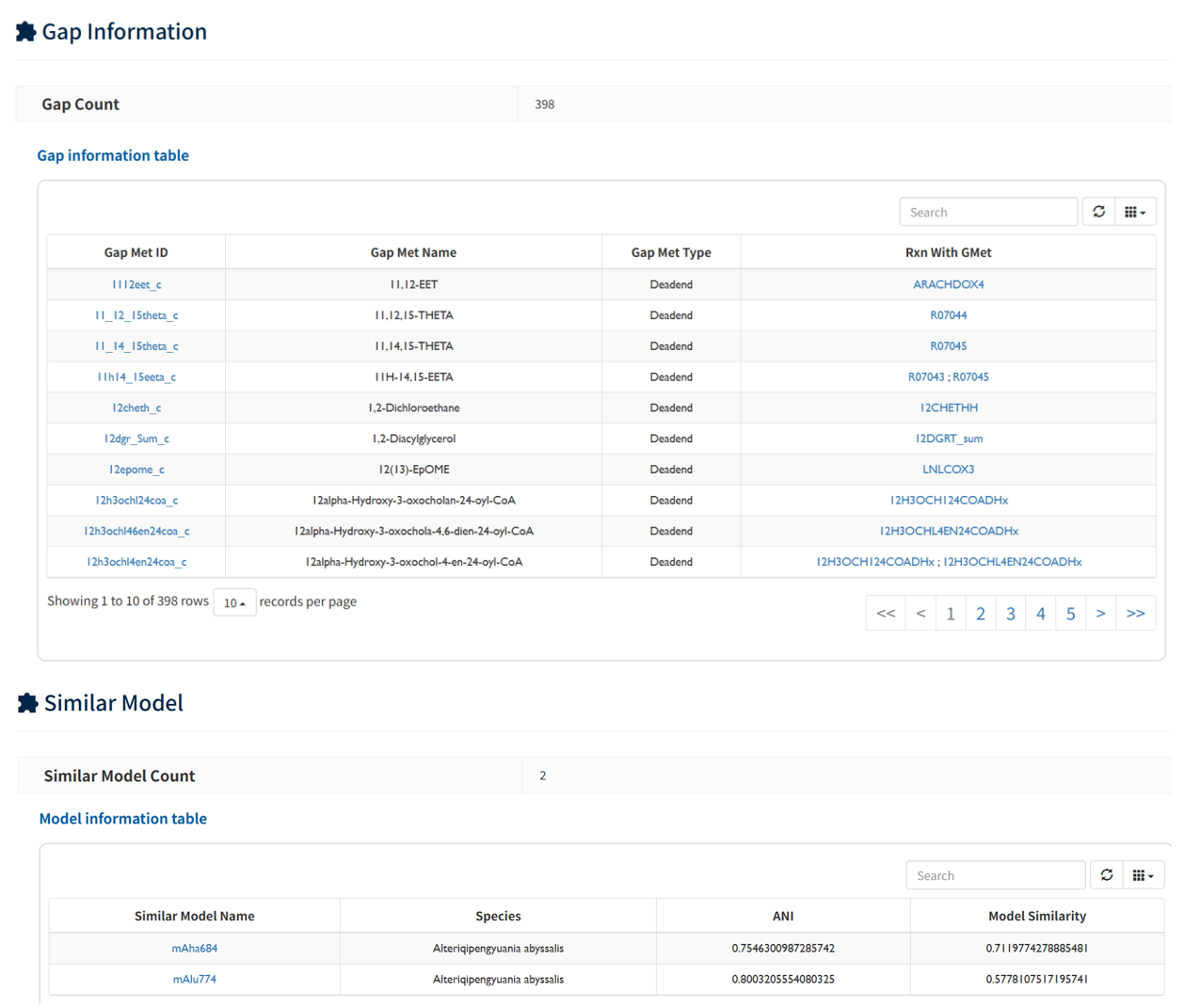
 In Gene/Reaction/Metabolite section, detailed annotations of model components, including genes, reactions, and metabolites, are presented in tabular format. In addition, gap-associated metabolites are provided to highlight potential missing links in the metabolic network. These detailed annotations facilitate a better understanding of the model structure . In Similar Model Section, similar models within the platform were listed, identified based on network topology, enabling users to explore and compare related metabolic systems.
In Gene/Reaction/Metabolite section, detailed annotations of model components, including genes, reactions, and metabolites, are presented in tabular format. In addition, gap-associated metabolites are provided to highlight potential missing links in the metabolic network. These detailed annotations facilitate a better understanding of the model structure . In Similar Model Section, similar models within the platform were listed, identified based on network topology, enabling users to explore and compare related metabolic systems.
Browse Reactions
The Browse reactions page provides all reactions within this platform. The name and equation information are present once you enter this page.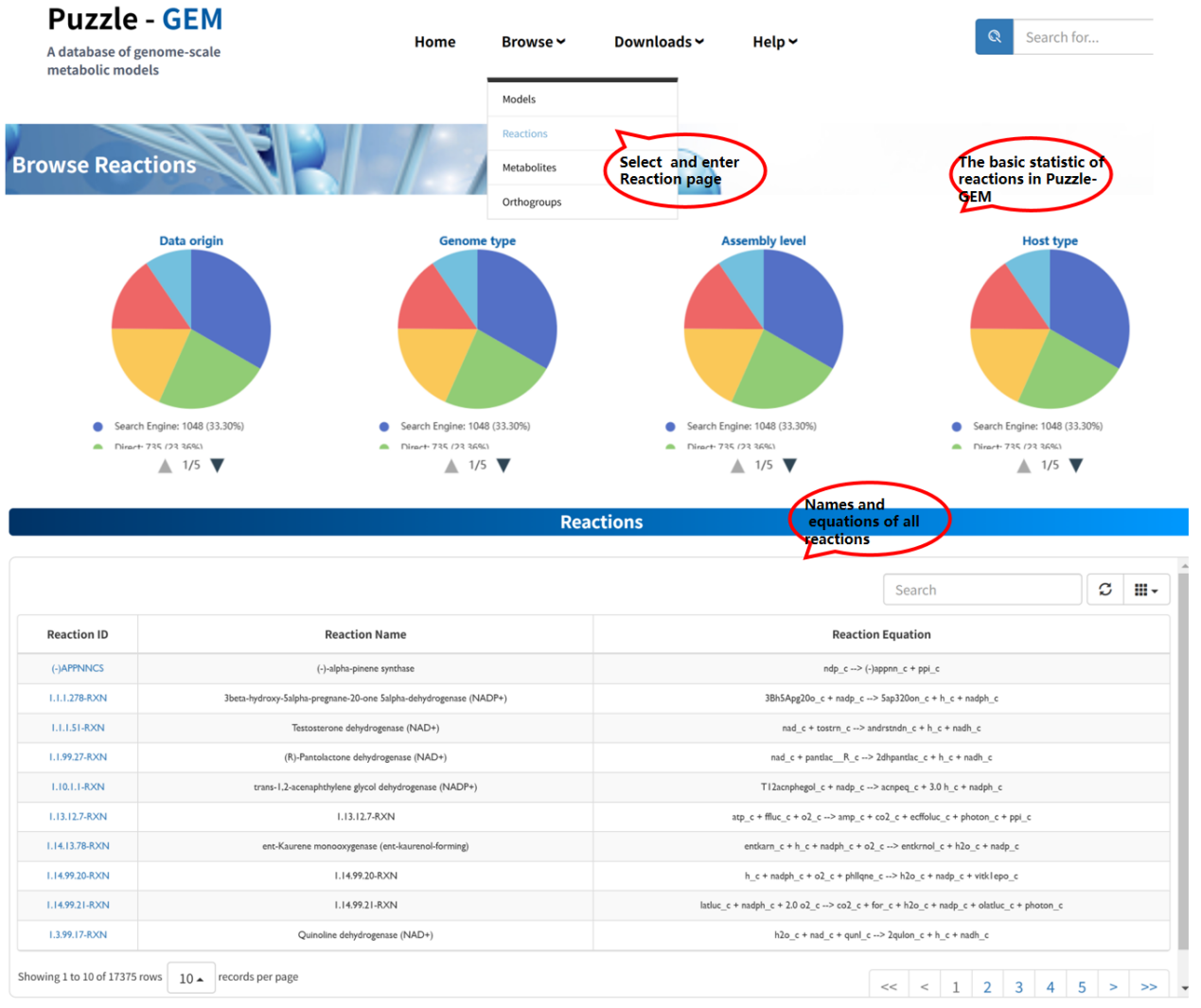 Users can access detailed information for each reaction by clicking the Reaction ID. Each reaction is associated with a dedicated Reaction information page, which consists of two sections: Basic Information and Models containing this reaction.
The Basic Information section summarizes the identifiers of the reaction across multiple databases, along with its fundamental properties, including reaction equation, reversibility, and functional annotations. In addition, the page displays the distribution of the reaction across species. A heatmap visualization summarizes its presence across different genera, organized according to phylogenetic relationships. The color intensity represents the number of species within each genus that contain the reaction, reflecting its evolutionary conservation and taxonomic distribution.
Users can access detailed information for each reaction by clicking the Reaction ID. Each reaction is associated with a dedicated Reaction information page, which consists of two sections: Basic Information and Models containing this reaction.
The Basic Information section summarizes the identifiers of the reaction across multiple databases, along with its fundamental properties, including reaction equation, reversibility, and functional annotations. In addition, the page displays the distribution of the reaction across species. A heatmap visualization summarizes its presence across different genera, organized according to phylogenetic relationships. The color intensity represents the number of species within each genus that contain the reaction, reflecting its evolutionary conservation and taxonomic distribution.
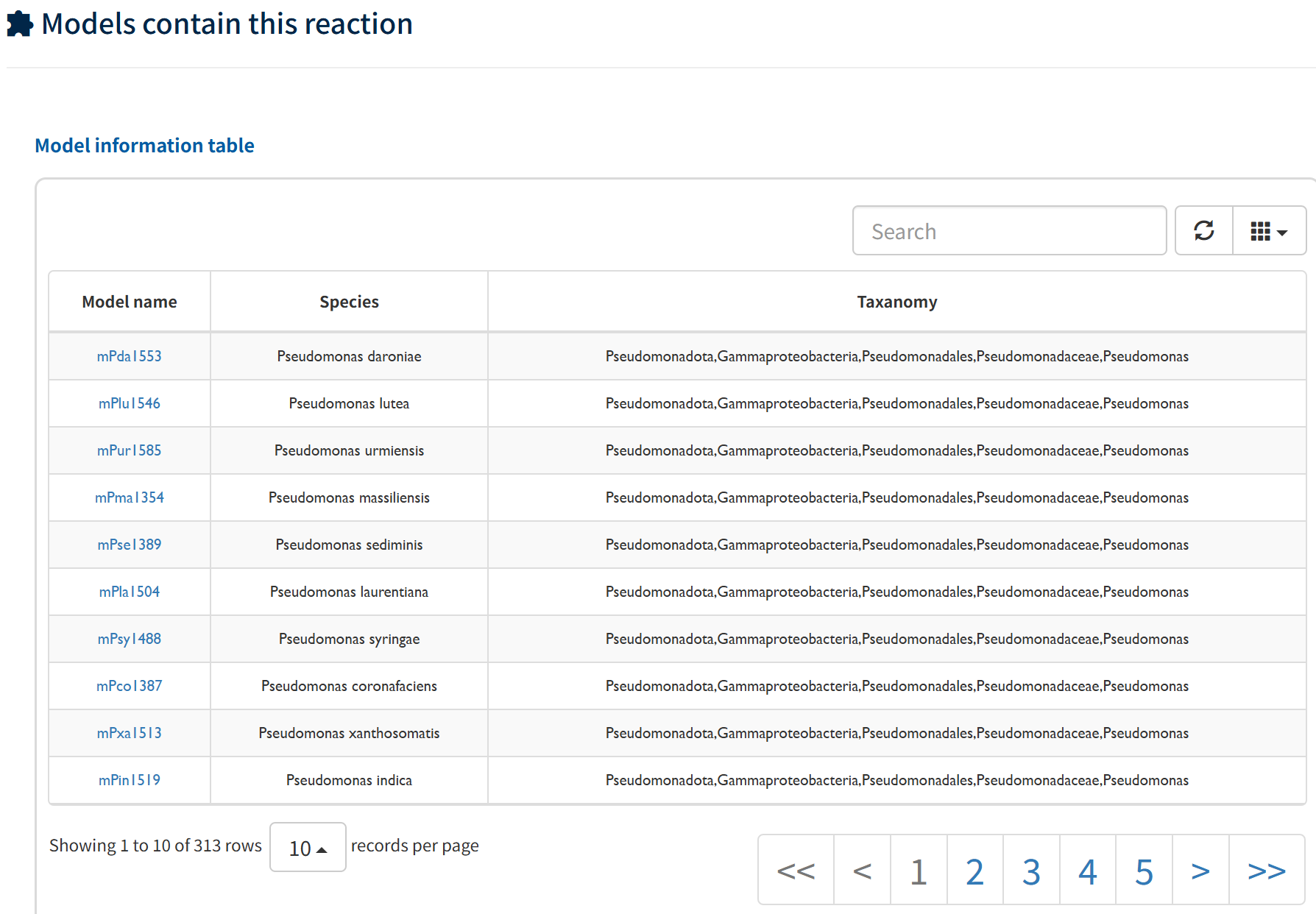
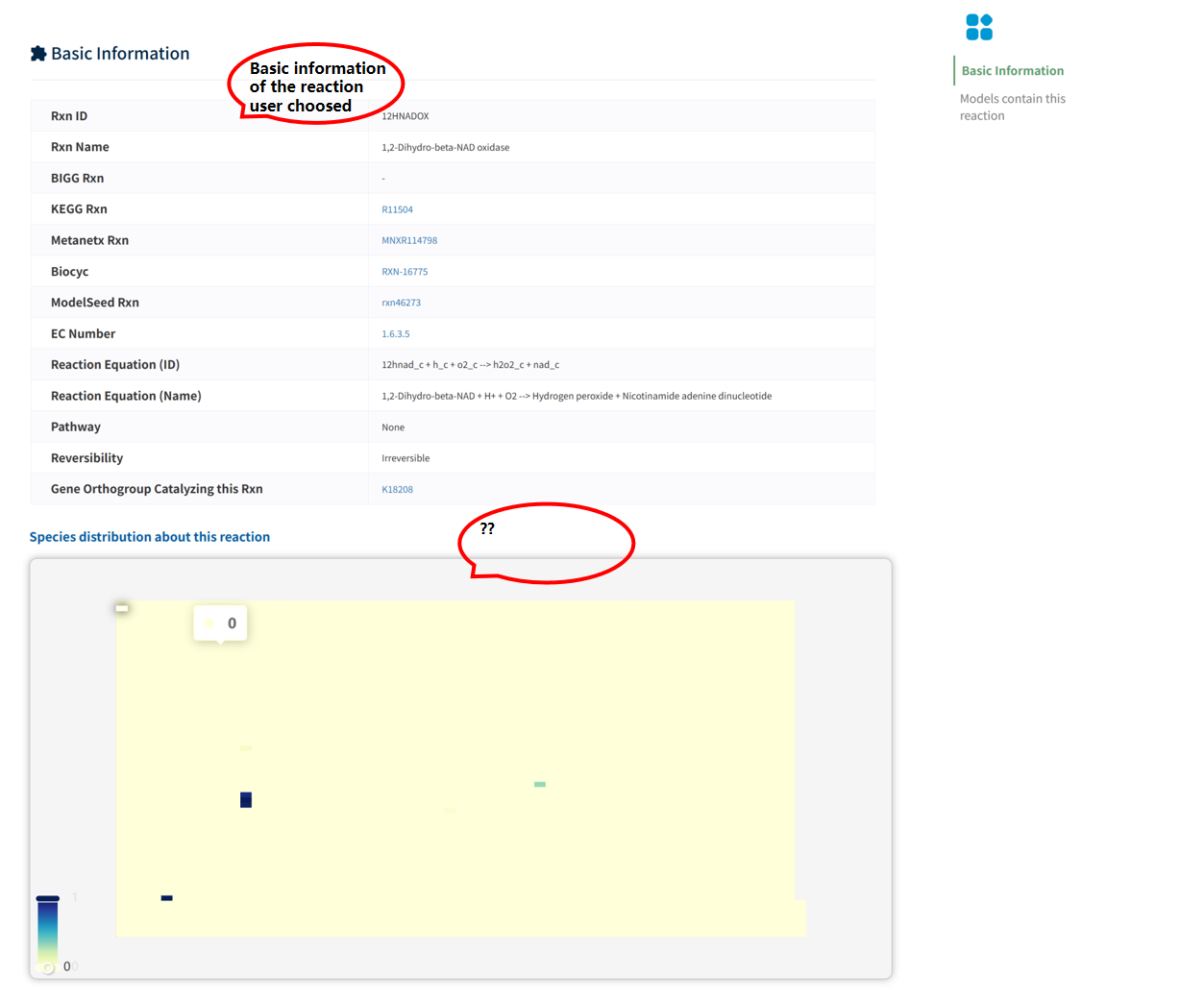 The Models containing this reaction section lists all metabolic models that include this reaction, along with their species and taxonomic information, enabling users to examine its distribution across different models and organisms.
The Models containing this reaction section lists all metabolic models that include this reaction, along with their species and taxonomic information, enabling users to examine its distribution across different models and organisms.
Browse Metabolites
The Browse metabolites page provides all metabolites within this platform. The name and formula information are present once you enter this page.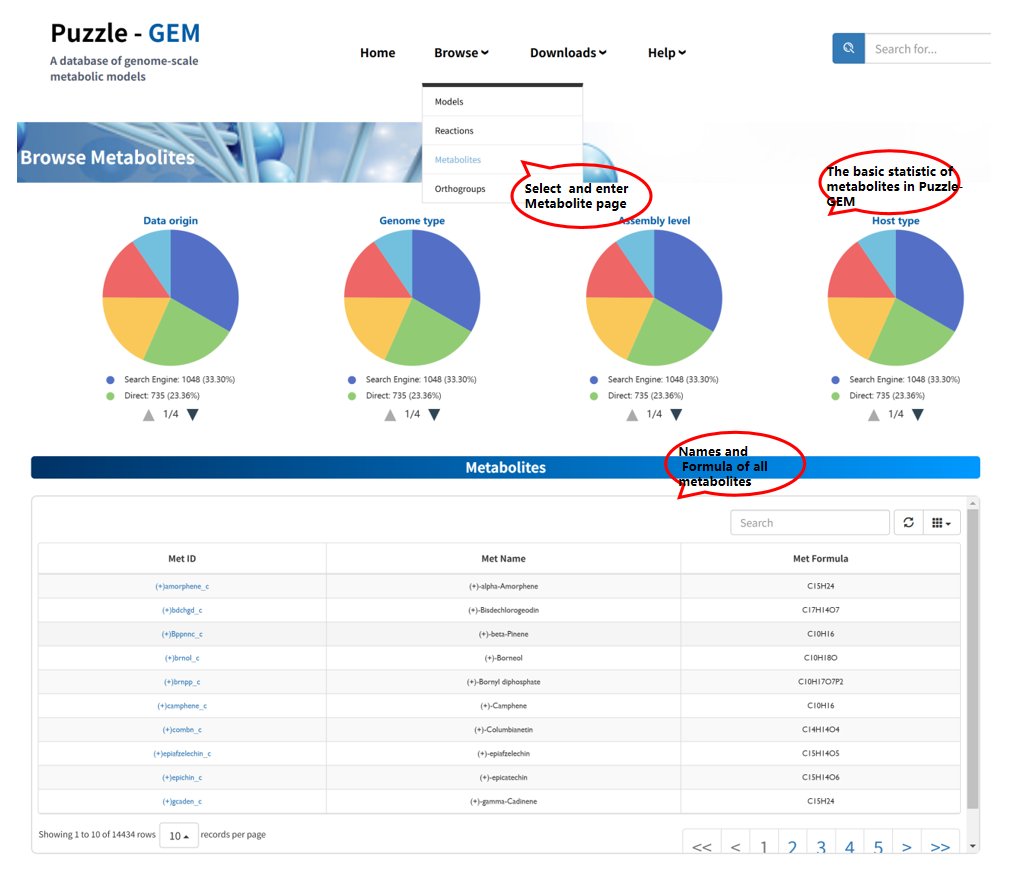 Users can access detailed information for each metabolite by clicking the Metabolite ID. Each metabolite is associated with a dedicated Metabolite information page, which consists of two sections: Basic Information and Reactions and Models containing this metabolite.
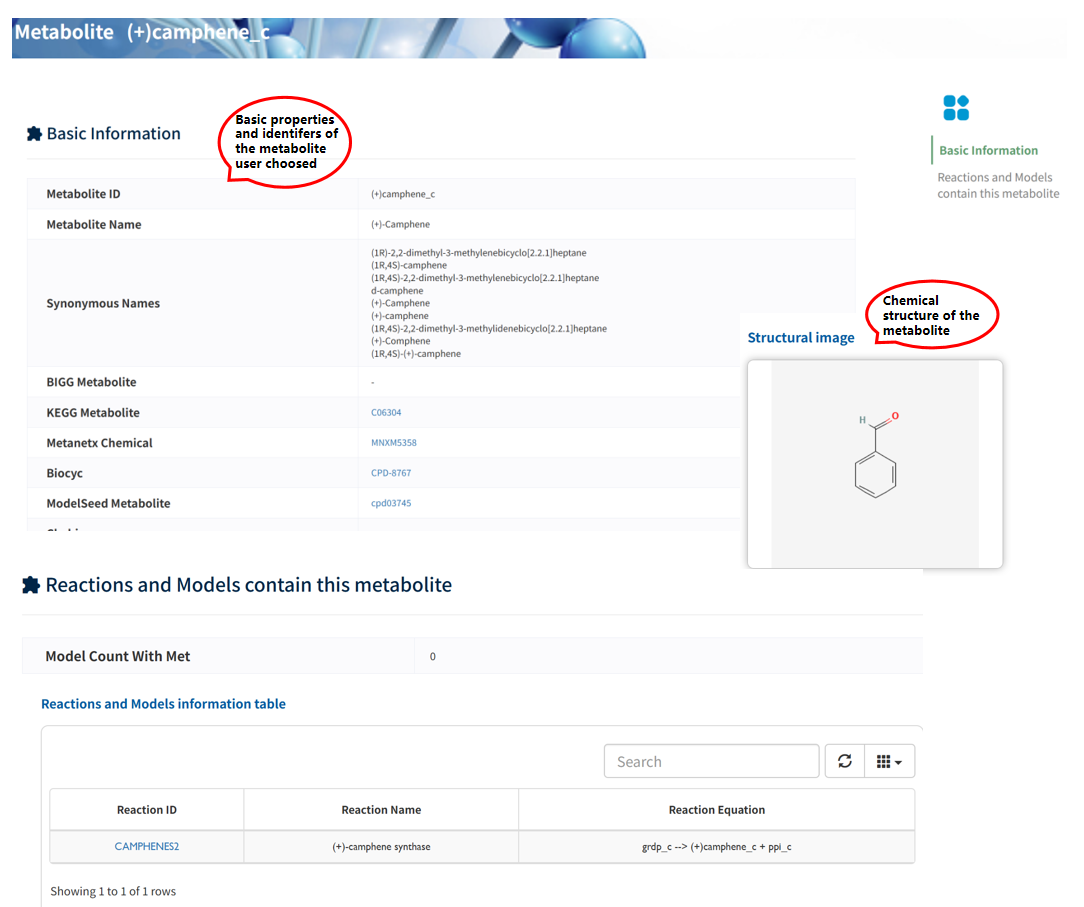
The Basic Information section summarizes the identifiers of the metabolite across multiple databases, along with its fundamental properties, including metabolite name, synonymous names, and cross-references to external resources (e.g., KEGG, MetaNetX, BiGG, and ModelSEED). In addition, a structural image is provided to illustrate the chemical structure of the metabolite, facilitating intuitive understanding of its molecular characteristics.
The Reactions and Models containing this metabolite section lists all reactions involving the metabolite, along with the associated metabolic models. This enables users to explore the functional role of the metabolite within metabolic networks and examine its occurrence across different models.
Users can access detailed information for each metabolite by clicking the Metabolite ID. Each metabolite is associated with a dedicated Metabolite information page, which consists of two sections: Basic Information and Reactions and Models containing this metabolite.
The Basic Information section summarizes the identifiers of the metabolite across multiple databases, along with its fundamental properties, including metabolite name, synonymous names, and cross-references to external resources (e.g., KEGG, MetaNetX, BiGG, and ModelSEED). In addition, a structural image is provided to illustrate the chemical structure of the metabolite, facilitating intuitive understanding of its molecular characteristics.
The Reactions and Models containing this metabolite section lists all reactions involving the metabolite, along with the associated metabolic models. This enables users to explore the functional role of the metabolite within metabolic networks and examine its occurrence across different models.
Browse Orthogroups
The Browse Orthogroups page provides all gene orthogoupsW within this platform. The function and basic statistics information are present once you enter this page.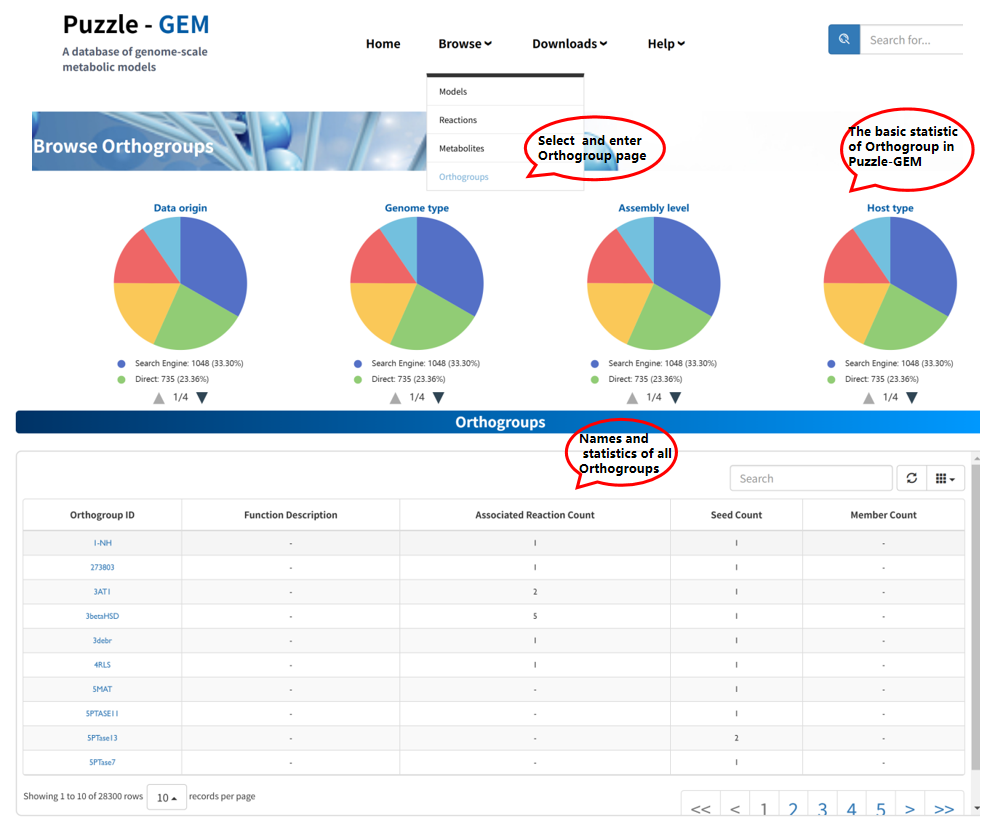 Users can access detailed information for each orthogroup by clicking the Orthogroup ID. Each orthogroup is associated with a dedicated Orthogroup information page, which consists of 3 sections: Basic Information, Seed Information, and Gene Member Information.
The Basic Information section summarizes the core attributes of the orthogroup, including its identifier, functional description, and associated reactions. This provides a direct link between gene orthology and metabolic function.
The Seed Information section lists the seed genes used to define the orthogroup, along with their species and data sources. This information helps trace the origin and reliability of the orthogroup construction.
The Gene Member Information section presents all gene members assigned to the orthogroup across different models, including their corresponding models and species.
Users can access detailed information for each orthogroup by clicking the Orthogroup ID. Each orthogroup is associated with a dedicated Orthogroup information page, which consists of 3 sections: Basic Information, Seed Information, and Gene Member Information.
The Basic Information section summarizes the core attributes of the orthogroup, including its identifier, functional description, and associated reactions. This provides a direct link between gene orthology and metabolic function.
The Seed Information section lists the seed genes used to define the orthogroup, along with their species and data sources. This information helps trace the origin and reliability of the orthogroup construction.
The Gene Member Information section presents all gene members assigned to the orthogroup across different models, including their corresponding models and species.
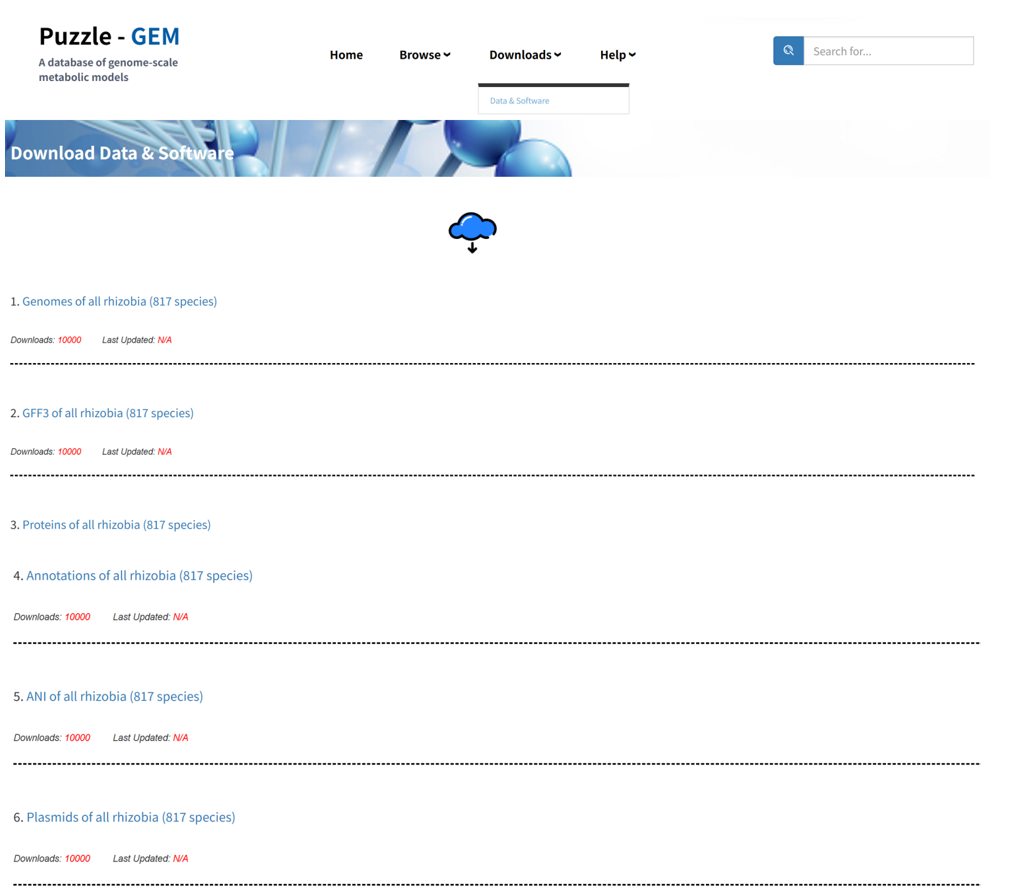
Downloads
Help
 Users can access detailed documentation of the entire platform by clicking the Tutorial page.
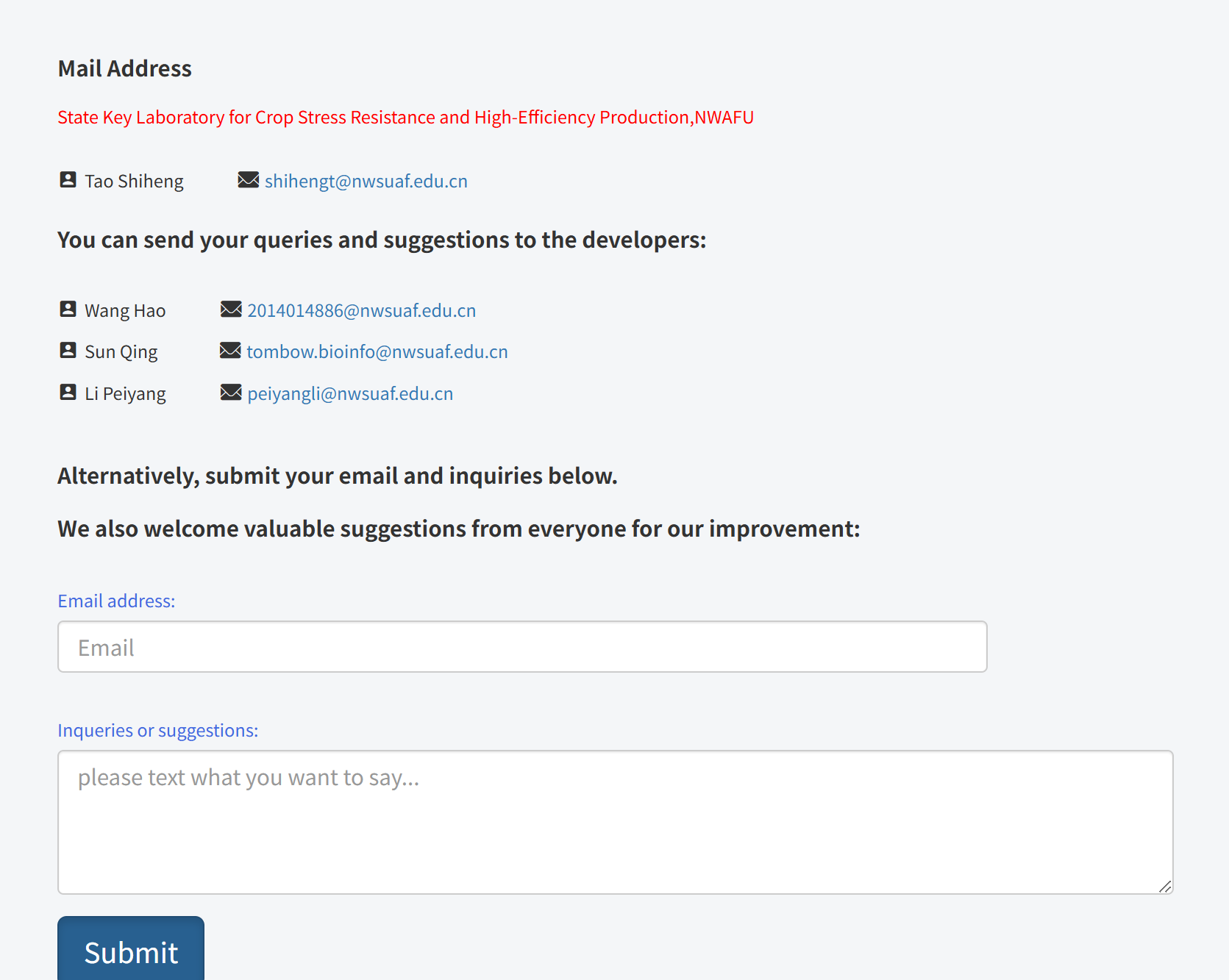
Additionally, in the Contact us page, users can send queries and suggestions to the developers via the provided email addresses or by submitting messages through the online form. This allows users to report issues, ask questions, and provide feedback for further improvement of the platform.
Users can access detailed documentation of the entire platform by clicking the Tutorial page.
Additionally, in the Contact us page, users can send queries and suggestions to the developers via the provided email addresses or by submitting messages through the online form. This allows users to report issues, ask questions, and provide feedback for further improvement of the platform.